DANA POINT, CA — Think of it as a physiological version of cloud computing, says functional neurologist, David Perlmutter, MD.
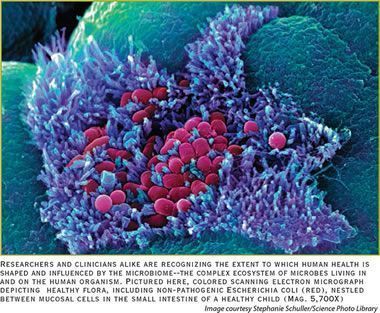 “The microbiome in the human gut represents over 3 million different genes. It is like a cloud computer for genes. We’ve put a lot of eggs in human genome basket, so it is humbling to realize that the human genome is only about 23,000 genes. Our lives and our health are hugely influenced by the organisms living in our gut,” Dr. Perlmutter said at the Nutrition Business Journal Summit.
“The microbiome in the human gut represents over 3 million different genes. It is like a cloud computer for genes. We’ve put a lot of eggs in human genome basket, so it is humbling to realize that the human genome is only about 23,000 genes. Our lives and our health are hugely influenced by the organisms living in our gut,” Dr. Perlmutter said at the Nutrition Business Journal Summit.
The microbiome—the diverse world of organisms living in and on the human body–is the new frontier in biomedical science, and it’s been a veritable bonanza for researchers over the last decade. What they are discovering has potential to radically reshape how we understand human health and how we treat illness.
Many seemingly disparate disorders have been linked to distinct changes in the type and proportion of different microbes living in the gut. Well beyond obvious GI conditions such as irritable bowel and colorectal cancer, researchers have also found connections between microbiome changes and depression, Alzheimer’s, autism, fibromyalgia, multiple sclerosis, even diabetes.
Microbiome research thus far has yielded equal parts good and bad news. On the positive side, it opens up possibilities for new, less invasive nutritional and probiotic-based approaches to preventing and treating so-called “incurable” chronic diseases.
On the downside, many aspects of modern life—from antibiotic and pesticide overuse to widespread cesarean sections—are seriously deranging the fragile yet essential ecosystems in our GI tracts, said Dr. Perlmutter who practices in Naples, FL, and is the editor in chief of a new journal called Brain and Gut, launching this month.
If the decoding of the human genome has been a game-changer in medicine, the mapping of the microbiome is creating an entirely different game.
Mapping the World Within
“It’s not just genomic differences (that influence health), but also microbiomics. Your ability to absorb something is dependent not just on your genome but also on your microbiome’s genome, and this is unexplored territory. We’re gathering all of that data, working with a lot of different populations. We’re especially interested in the ‘idiopathically ill,'” said Jessica Richman, CEO and co-founder of μBiome, a San Francisco start-up hatched at the University of California San Francisco’s Quantitative Bioscience Institute.
In 2013, the company raised over $351,000 from roughly 2,500 donors on indiegogo.com. It is one of the most successful crowdfunding campaigns to date. μBiome has also raised large sums in start-up capital from angel and venture investors. The company’s goal? “To equip all individuals with the tools they need in order to empower them to learn about the unique balance of bacteria in their bodies.”
μBiome’s jump-off point is the National Institutes of Health’s $115 million Human Microbiome Project, an ongoing endeavor begun in 2007 that has gene-sequenced entire microbiomes from 242 healthy but diverse humans so far.
The NIH project has identified roughly 10,000 unique microbial species living in or on the study subjects. Any one individual plays host to between several hundred to several thousand of these commensal species.
Under the motto, “Big data from bacteria,” μBiome takes microbiome research out of the Ivory Tower and into the street, offering an $89 kit enabling people to learn exactly what bacterial species live in and on their bodies, in what proportion, and how their microbial universes compare with those of demographically similar people.
Big Data from Bacteria
The basic kit covers the GI microbiome—a user simply swabs a fecal sample from toilet paper and sends it to μBiome for analysis. For additional fees, users can also sample the nose, ear and urogenital tract. μBiome is utilizing the state of the art Illumina sequencing platform to analyze the microbial DNA.
To date, the company has analyzed samples from several thousand people all over the world. One of the company’s main goals is to determine how much a given individual’s microbiome composition is influenced by the human host’s genetics, and how much by diet & lifestyle factors.
μBiome positions itself as an open-source “citizen science” research project. It provides no specific diagnostics or health assessments. Though it does it does invite users to track their microbiomes as they make diet and lifestyle changes, μBiome is clearly trying to steer a course away from classification as a diagnostic device.
Bacteroides vs Firmicutes
Nearly all of the bacteria in the gut–roughly 99%–are anaerobes. Worldwide over 35,000 unique microbial species have been identified (so far) in humans, and the vast majority fall into two main phyla: Bacteroides and Firmicutes.
Bacteroides species are particularly important because they play a critical role in breaking down and transforming undigestible carbohydrates from plant foods into short chain fatty acids and other absorbable nutrients. In essence, they are digestive and metabolic helpers that greatly increase the nutritive value of plant-based foods.
The relative proportion of Bacteroides to Firmicutes in someone’s gut is an important factor in overall health, said Dr. Perlmutter.
In a landmark study several years ago, Carlotta De Filippo and colleagues of the University of Florence compared the gut microflora in healthy children in a rural village in Burkina Faso, West Africa with those of similarly aged kids Italian kids living in Florence.
The West African kids, from the Mossi ethnic group, typically ate high-fiber diets rich in complex carbs, vegetables and legumes, and low in animal protein. Stone-ground millet and sorghum are staples in Burkina Faso, often eaten as a porridge. The European kids, ate much higher amounts of animal protein and animal fat as well as vastly more refined carbs, sugar, artificial sweeteners, and additives.
Overall, Mossi children showed a greater diversity of gut microflora than the Italian kids. In general, microbial diversity is thought to be a sign of a healthy microbiome; a paucity in microbial variety predisposes to illness.
The African kids showed a much larger proportion of Bacteroides to Firmicutes than the Europeans. Bacteroides species represented 58% of the total microbiome in the Africans, versus 22.4% in the Europeans. In contrast, Firmicutes comprised 64% of the microbiome in the Italian kids versus just 27% in the Mossi group. “The differential distribution of Firmicutes and Bacteroidetes delineates profound differences between the two groups,” the authors note (De Filippo C, et al. PNAS. 2010; 107(33)).
The Burkina Faso children also had high prevalence of Prevotella and Xylanibacter, organisms that were completely absent in the European kids. These are capable of cellulose and xylan hydrolysis.
There was a significantly higher level of short-chain fatty acids in the fecal samples from the African versus the European kids, with markedly higher levels of acetic and butyric acids. In contrast, the Italians had higher levels of propionic acid.
This has important implications, said Dr. Perlmutter. Propionic acid alters neurotransmitters and immune function, and reduces glutathione, increases Omega-6 levels, and in animal experiments has been shown to have significant negative effects on brain function.
In summarizing the comparisons, Dr. De Filippo wrote, “gut microbiota co-evolved with the polysaccharide-rich diet of Burkina Faso individuals, allowing them to maximize energy intake from fibers while also protecting them from inflammation and non-infectious colonic diseases.” She noted that potentially pathogenic bacteria like Shigella, Salmonella and Escherichia were more common in the Italian than the African children.
She attributed the relative lack of microbial diversity in European children to high consumption of sugar, animal fat and calorie-dense processed foods that may favor overgrowth of less beneficial species. The “microbial simplification” observed in the Italian kids, “harbors the risk of depriving our microbial gene pool of potentially useful gene reservoirs.”
A Role in Cognitive Function
According to Dr. Perlmutter, the gut microflora play a major role in neurocognitive conditions like autism. “An autistic brain is an inflamed brain, and autistic kids have a very characteristic microbiome pattern. It’s almost a fingerprint,” he told the NBJ Summit.
Most children with autism spectrum disorders have histories of significant antibotic exposures, chronic gastrointestinal problems, and abnormal food cravings. It turns out that they are likely to have an overload of Clostridium in their GI tracts.
Parracho and colleagues at the University of Reading, UK, studied the microbiomes of 58 autistic kids, as well as 12 non-autistic sibilings, and 10 non-autistic and unrelated children of similar age.
The most striking finding was a preponderance of Clostridium histolyticum in the autistic kids compared with their non-autistic siblings or unrelated peers (Parracho HM, et al. J Med Microbiol. 2005; 54(10):987-991).
“Clostridia are recognized toxin-producers, including neurotoxins. Theoretically, toxic products may be over-expressed in the autistic gut, which may lead to increased levels in the bloodstream and thus exert systemic effects. Interestingly, many anecdotal reports from parents of autistic children report worsening of behavioural symptoms coinciding with bouts of GI problems,” the Reading authors note.
Reflecting on the findings, Dr. Perlmutter pointed out that “If you treat autistic kids with vancomycin, used to treat C. difficile infections, you see a dramatic improvement in their behavior and attention.” This is not to suggest that vancomycin be used to treat autism; it simply illustrates the point that the microbiome changes may play a far greater role in cognitive function than previously recognized.
Belly Bugs & BDNF
Circulating levels of brain-derived neurotrophic factor (BDNF) is proving to be an important predictor of the risk of dementia, and gut bacteria may play a role in triggering production of this substance. Those in the lowest BDNF quintile had nearly twice the risk as those in the highest quintile.
Data from 2,131 dementia-free older adults (60-plus years) in the Framingham Heart Study showed a clear correlation between levels of circulating BDNF and risk of developing dementia within 10 years (Weinstein G, et al. JAMA-Neurology. 2014: 71 (1): 55-61).
Exercise, sunlight, Omega-3s (especially DHA), and some of the SSRI drugs have been shown to raise BDNF in humans. There is some compelling animal research suggesting that use of a Bifidobacterium longum supplement can also raise BDNF. Lab animals raised in sterile conditions and deprived of exposure to normal gut flora show markedly depressed levels of hippocampal BDNF production. This can be reversed by giving them B. longum.
“This is still experimental, but it is where we’re going in the future,” said Dr. Perlmutter.
Another recent study showed that a fermented milk product containing Bifidobacterium lactis, Lactobacillus bulgaricum, and several other probiotic species could reliably modulate midbrain emotional reactivity as measured by functional MRI, in a cohort of healthy women.
Compared with women taking a non-probiotic control, those who used the fermented milk product for 4 weeks showed brain changes suggestive of greater calm and relaxation during emotional processing tasks (viewing human faces & matching the expressions to specific emotions).
“Four-week intake of a fermented milk product with probiotics by healthy women affected activity of brain regions that control central processing of emotion and sensation,” concluded author Kirsten Tillisch, of the Geffen School of Medicine, University of California-Los Angeles (Tillisch K, et al. Gastroenterology. 2013; 144 (7): 1394-1401).
The microbial world within the lumen of the digestive tract profoundly affects glucose and fat metabolism, and a growing number of studies suggest that the microflora are a major factor in obesity.
Influence on Obesity
A number of studies from research groups worldwide indicate that the gut microbiome strongly influences propensity toward weight gain, and may therefore play a role in obesity.
In a 2012 review article on this emerging topic, Krznaric and colleagues underscored the apparent link between obesity and a decrease in the relative proportion of Bacterioides to Firmicute species in the gut.
Investigators at the National Research Center in Cairo, Egypt studied the gut microbial populations in a cohort of 79 children and adults, 51 of whom were obese. They noted that the proportion of Firmicutes to Bacteroides was distinctly different in the obese individuals, skewing toward the Firmicutes.
The obese individuals showed higher levels of Firmicutes as well as hydrogen-producing Bacteroides species. The authors speculated that the presence of these organisms, “accelerate the fermentation of otherwise indigestible carbohydrates, thereby leading to increased acetate production and subsequently higher energy absorption.”
They also found that this change was associated with higher levels of C-reactive protein, an important indicator of systemic inflammation (Ismail NA, et al. Arch Med Sci. Jun 2011; 7(3): 501–507).
Research on the role of the microbiome in obesity is still in a very preliminary stage, but the existing body of knowledge does have clinical implications: childhood obesity correlates with antibiotic exposure.
In a 2012 study conducted at New York University, children who had been given antibiotics before the age of 6 months were 22% more likely to become overweight by the time they entered preschool than kids who had never been exposed to these drugs. This human data echoed an earlier rodent study showing weight gain and increased fat storage following antibiotic exposure.
A Role in Cancer
Investigators are just beginning to explore the potential impact of the microbiome on cancer risk. It is reasonable to expect that it would play a role, at least in cancers of the gastrointestinal tract.
This summer, University of Michigan researchers reported that analysis of the microbiome is more effective in predicting the presence of precancerous adenomatous polyps and invasive colorectal tumors than fecal occult blood testing.
Dr. Patrick Schloss and colleagues at the University’s Department of Microbiology & Immunology analyzed stool samples from 90 people: 30 healthy individuals, 30 patients with precancerous adenomatous polyps, and 30 patients with invasive colorectal cancer. They found that the composition of the gut microbiome was distinct for individuals in the three groups.
Using this information, they then clarified microbiome signatures for each group, and found that people with precancerous or cancerous lesions had marked depletions in a number of commensals found in the gut of the normal cancer-free subjects.
Dr. Schloss’ team then developed a risk assessment tool based on the microbiome data, and found that by looking for the “signature” microbial changes, they obtained a 4.5-fold improvement in the detection of adenomatous polyps, and a 5.4-fold improvement in the detection of invasive colorectal cancer (Zackular JP, et al. Cancer Prev Res; epub ahead of print)
“Gut microbiome analysis has the potential to be a new tool to noninvasively screen for colorectal cancer,” said Dr. Schloss.
END