Just when you thought the microbiome couldn’t get any weirder, investigators at Stanford University have discovered a whole new dimension in the microbial universe: a category of distinct, self-replicating RNA sequences in and among bacteria of the human gut and oral microbiomes.
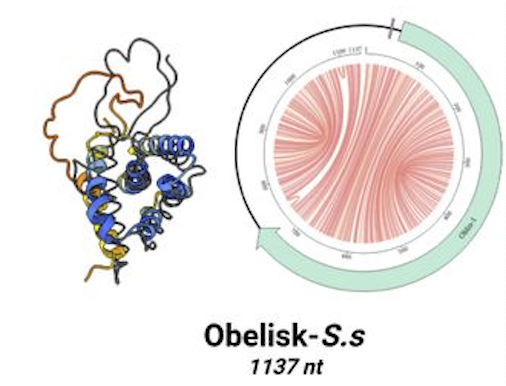
Named “Obelisks” owing to their circular rod-like structures, these peculiar bits of genetic information are unlike any genomic sequences in the known viral, bacterial, fungal, or animal worlds.
In discovering Obelisks, Nobel prize-winning geneticist Andrew Fire and his team believe they’ve opened an entirely new frontier in microbiology.
“Obelisks form their own distinct phylogenetic group with no detectable sequence or structural similarity to known biological agents,” they write in their newly published article on bioRxiv.com, the Cold Spring Harbor Laboratory’s preprint server platform.
A New Frontier
So far, Fire and his colleagues have identified almost 30,000 distinct Obelisks, by applying a bioinformatics tool called VNom (Viroid Nominator) to databases containing 5.4 million genome sequence datasets.
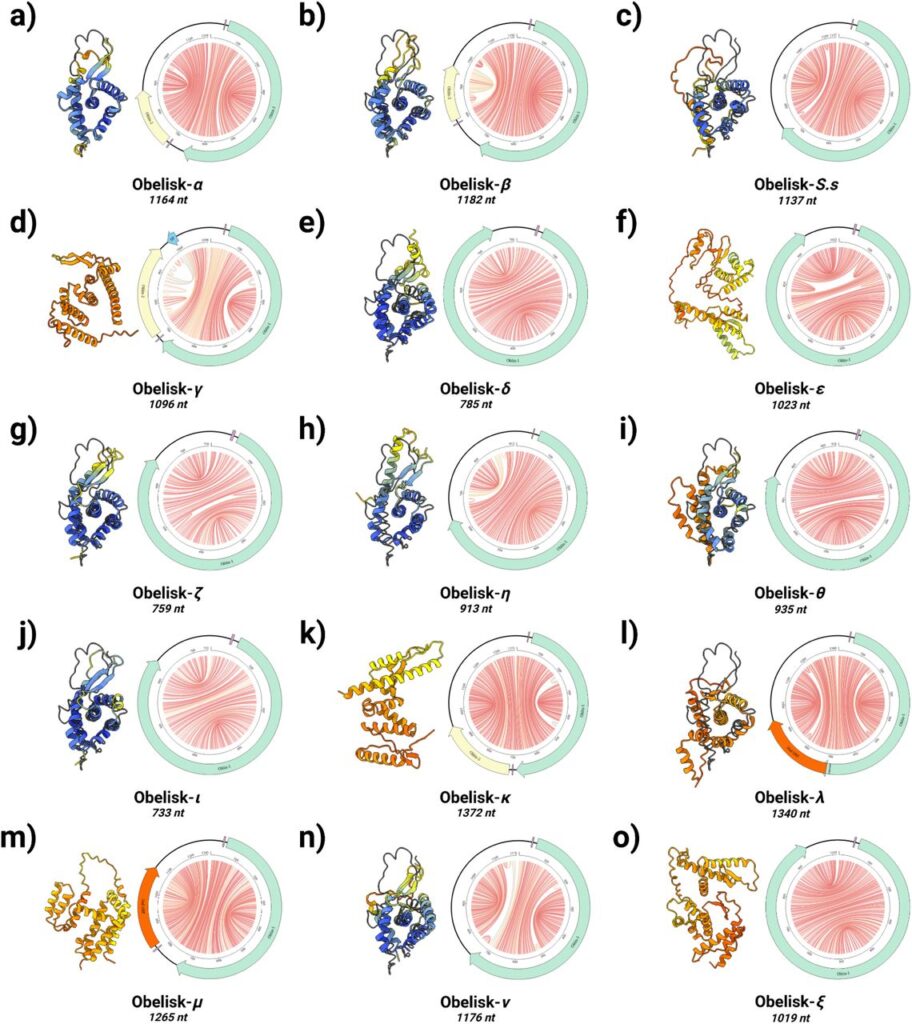
Obelisks are diverse in structure, but they share several common features. They’re all circular RNA strands of no more than 1kb in length. That’s tiny even by the miniscule standards of molecular biology. They form predictable rod-like secondary structures. And they contain open reading frames that code for unique, previously unknown protein-like entities that the Stanford group has named “Oblins.”
In analyzing microbiome data sets representing the oral and gut microbiomes of 472 healthy human donors, they found that nearly 10% of these individuals carried one or more Obelisks among their commensal bacteria. These curious RNA sequences were more prevalent in oral versus gut microbiome samples (53% versus 6.6%), which suggests that the oral flora may be a reservoir for Obelisks in humans.
Streptococcus sanguinis, a facultative anaerobe that is part of the normal oral flora, is the first bacterium within the human microbiome that the Stanford team has identified as a definite host for Obelisks. The specific Obelisk subtype that Fire and colleagues discovered inside S. sanguinis, is able to replicate, and to produce a certain type of Oblin within the Streptococcal cells.
“Obelisks form their own distinct phylogenetic group with no detectable sequence or structural similarity to known biological agents.”
–Andrew Fire, Professor of Molecular and Genetic Medicine, Stanford University
This, they say, is clear evidence that Obelisks can and do live—if such entities can be considered living–within bacterial cells. Though other bacterial hosts for Obelisks have not yet been identified, Fire says “it is reasonable to assume that at least a fraction (of the Obelisks) may be present in bacteria.”
Obelisks are Everywhere
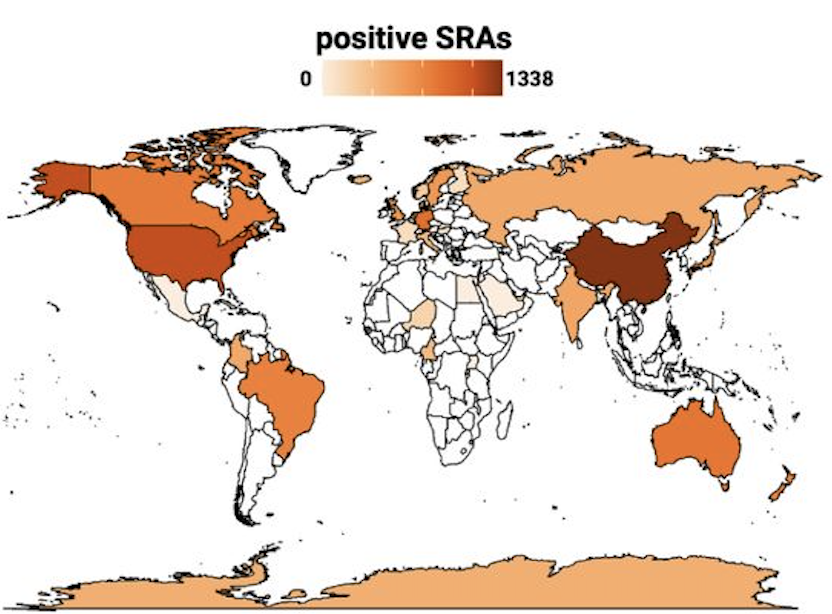
Inside or outside of bacterial host cells, Obelisks seem to be everywhere. The Stanford group has detected them, at varying prevalence levels, in metatranscriptomes from people living in the US, Canada, Brasil, China, India, Russia, and Australia, as well as several countries in Europe and Africa. They’ve also found them in non-human mammals, fish, birds, arthropods, even in soil and sediment. Wherever there is life of any sort, there appear to be Obelisks.
Are Obelisks friendly? Fearsome? Both? At this point, nobody knows.
They do share some structural traits with viroids—the small, non-encapsulated DNA or RNA strands typically found in plants, and which are sometimes pathogenic. The potato spindle tuber viroid (PSTVd), and the citrus exocortis viroid (CEVd) are two common examples of pathogenic plant viroids.
Like viroids, Obelisks are very small circular RNA sequences that lack protein coats. This absence of a protective protein capsule is the key feature that distinguishes both viroids and Obelisks from true viruses.
So far, none of the 29,959 Obelisks discovered by Fire and his team appear to cause harm to the bacteria among which they’re found, or to their human meta-hosts. And there’s not yet enough information about them to determine whether they are helpful.
Are Obelisks friendly? Fearsome? Both? At this point, nobody knows.
It is clear that Obelisk RNA sequences can be transcribed into protein-like structures, but the function of these so-called Oblins is an open question.
The Stanford researchers have found only one such Oblin that has an identifiable function: a self-cleaving ribozyme similar to the hammerhead type-III ribozymes. Hammerhead ribozymes are RNA catalytic cleavage sequences found in genomes throughout the bacterial, viral, and eukaryotic realms.
Beyond that, almost everything else about Obelisks and Oblins is a mystery at this point.
A Cornucopia of Questions
Lead investigator Andrew Fire is no stranger to the frontiers of knowledge. In 2006, he and his colleague Craig Mello won a Nobel Prize in Physiology or Medicine for their discovery of RNA interference (RNAi)—a mechanism that regulates gene expression. At the time, their elucidation of RNAi revolutionized scientific understanding of how genomes self-regulate. Their work underscored the ways in which tiny bits of double-stranded RNA can turn off gene transcription.

In the years since, Fire and his group have continued to delve into the mysteries of RNA, and they’ve developed sophisticated methods for detecting previously unidentified RNA sequences. It is this work which led to the discovery of Obelisks.
In his new paper, which has yet to be peer-reviewed, Fire acknowledges that the discovery of RNA Obelisks raises a cornucopia of interesting questions: How do Obelisks replicate? How do they move from extra- to intracellular environments? How many different Obelisk subtypes are there and what do they code for? What types of bacteria are most or least amenable to hosting Obelisks? Do the Oblins transcribed from the Obelisks have any important biological functions?
In microbiome data sets representing the oral and gut microbiomes of 472 healthy human donors, nearly 10% of these individuals carried one or more Obelisks among their commensal bacteria.
There are also big questions about how Obelisks affect the microflora in and around which they dwell, and by extension, the higher organisms that host these commensal microbes.
“Are Obelisks plasmid-like in that they can co-exist, and in some cases, contribute to host adaptability and fitness?” Fire asks. “Are they largely a deleterious or beneficial element to harbour? And what impact, if any, does harbouring an Obelisk have on ‘meta’-host physiology? Is Obelisk positivity predictive of human health states?”
Again, the answer to all of these questions is: We don’t yet know.
A New ‘ome?
It is notable that Obelisk prevalence, like microbiome composition, varies by anatomic site: the presence of Obelisks is much higher in the oral versus the intestinal microbiome.
Obelisks may—at least in part—be “colonists” within specific types of bacteria, in a similar way that various commensal bacteria have specific preferences for colonizing particular organ systems.
By way of rough analogy, the Obelisks might be something akin to the microbiome’s microbiome.
Dr. Fire speculates that if that proves to be the situation, then “donor-specific factors such as diet or lifestyles therefore likely play a role in Obelisk (re-)colonisation and retention.”
He adds that further study of S. sanguinis will be an important line of research in clarifying the relationship between these newly discovered RNA agents, the various microbiomes, and human physiology.
“Study of the implied S. sanguinis-Obelisk relationship might begin to reveal the relevance of Obelisks to the natural oral niche and potentially to human health, as well as offer a tractable model system to study Obelisk molecular biology.”
On the Threshold of Life
Researchers now know vastly more about the interrelationship between commensal microbes and human physiology than they did 20 years ago. But the Stanford group’s discovery of viroid Obelisks underscores just how much there is yet to discover about the invisible worlds that shape our existence.
This is especially true of viruses and related entities. In fact, the virome may be even more complex, influential, and downright weird than the bacterial microbiome.
According to an article by UC San Diego’s David Pride, and Rockefeller University’s Chandrabali Ghose, there are an estimated 380 trillion viruses inside the average human body. That’s ten times the already staggering number of bacteria in the typical human microbiome.
The overwhelming majority of viruses and viroid particles appear to be harmless. Those that are harmful are usually pathogenic to specific species. There are very few examples of viruses that cause disease throughout the animal kingdom.
Some viruses may actually be helpful. Researchers now know that genes from viruses known as HERVs (human endogenous retroviruses) account for up to 9% of the total human genome.
Obelisks appear to dwell at the threshold between entropy and order, between the realm of living things and the vast universe of inanimate matter. Their discovery is a brilliant burst of light into a domain that researchers have termed “viral dark matter.”
So far, early responses to the discovery from microbiologists and molecular biologists have been enthusiastic and positive. Beatriz Navarro, a plant viroid expert based at the Institute for Sustainable Plant Protection in Bari, Italy, praised it as “impressive,” in an interview published in New Scientist.
“Really wild and cool discovery for virus nerds,” gushed Steven Salzberg, a biostatistician at Johns Hopkins University.
Computational biologist Simon Roux, at the DOE Joint Genome Institute at Lawrence Berkeley National Laboratory, told the journal Science that he is “really impressed by the approach” that Fire and colleagues took in their work.
Roux is among a group of scientists who surmised that viroids—or similar non-encapsulated entities—might exist beyond the botanical world, inside commensal bacteria. The Stanford discoveries give credence to that hypothesis.
He added that the discovery of Obelisks begs the important question of whether true viruses evolved out of things like Obelisks and viroids, or rather the other way around–that Obelisks and viroids are the result of de-evolution of viruses or other more complex forms of life.
It will probably be years before we know if Obelisks have a role in human health or illness. But it is clear that their discovery is the first step in what will likely be a long and exciting scientific odyssey.
END